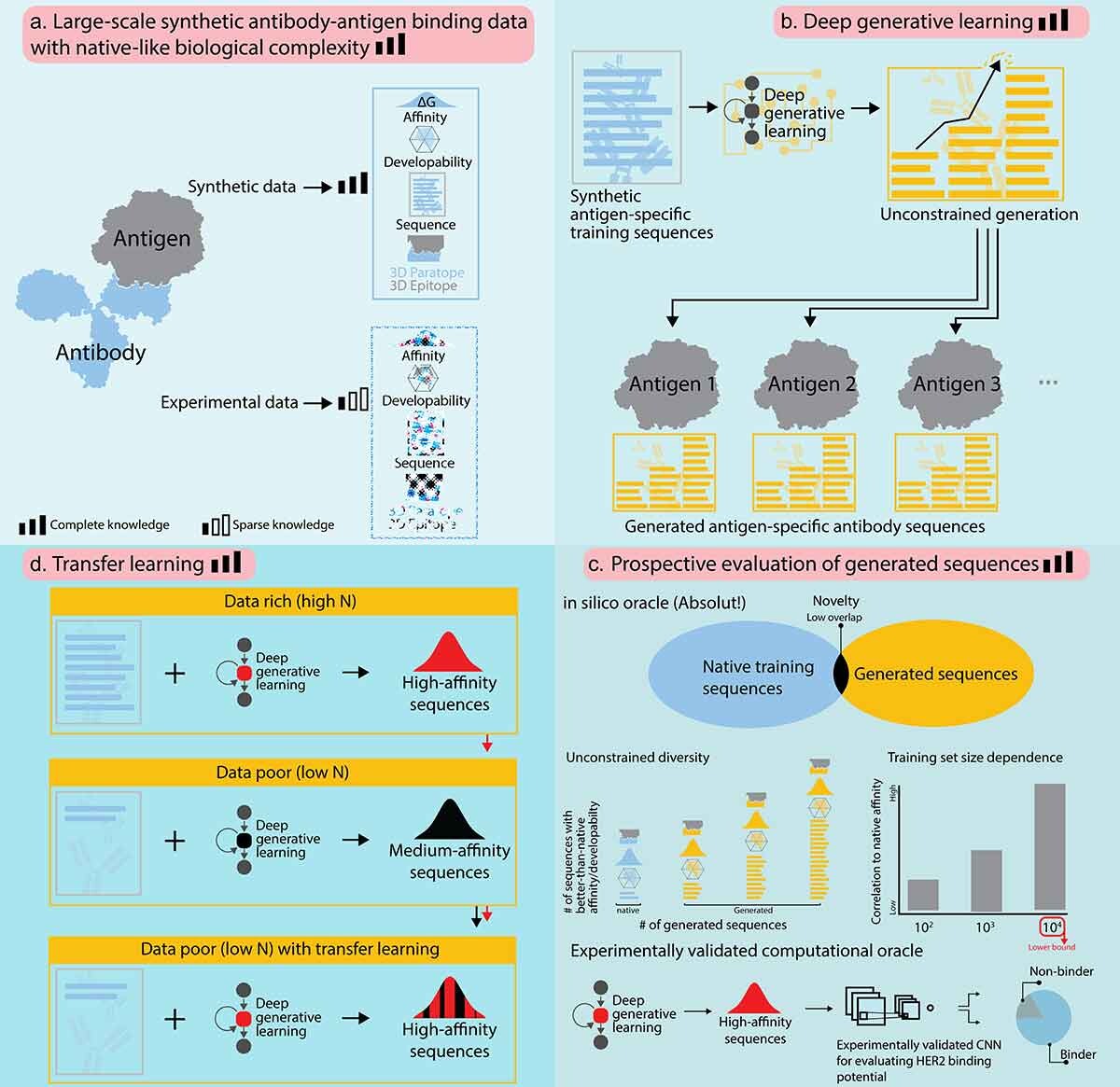
1) Do generative ML models keep the binding diversity of training antibodies?
Question: If I train a generative ML model with 10 000 antibodies that bind different regions (epitopes) on the antigen, can it generate equally diverse binders and discover novel sequences?
What did the ML learn, and can the ML generate antibodies as diverse, i.e., that bind all those epitopes. Can the model actually find new sequences?
Approach: We generate synthetic high-affinity sequences and train a generative model without exposing structural labels. Generated sequences are then tested back in Absolut!.
Indeed, these sequences keep the binding diversity of the training dataset.
Conclusion: The generated sequences preserve binding diversity without knowing structural information.The ML model also found many novel high-affinity candidates.
See full paper published in mAbs:
In silico proof of principle of machine learning-based antibody design at unconstrained scale
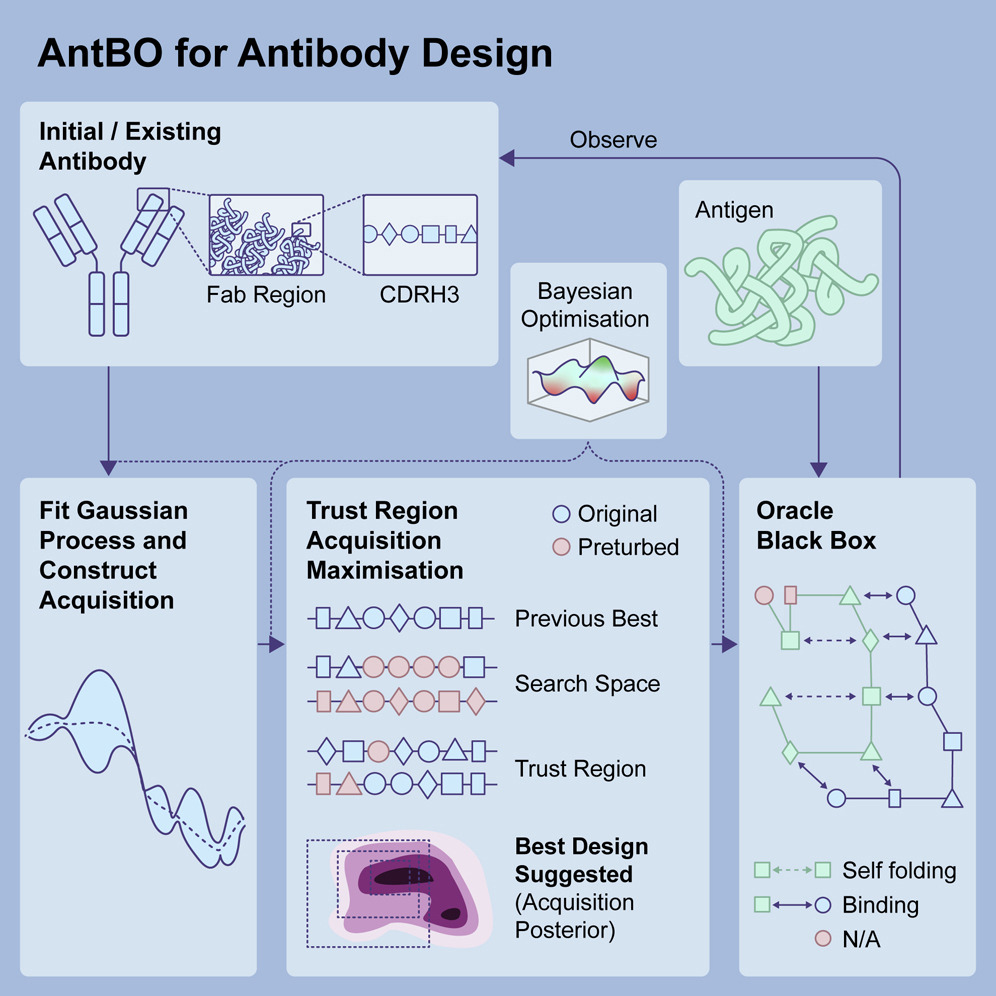
2) Minimal experimental dataset for reinforcement learning-driven lead optimization
Question: What is the minimal number of affinity measurements needed to create high-affinity antibodies to the same target?
Idea: Use adaptive ML: start with 10 antibodies → train → request 10 more → iterate. Bayesian ML predictions are validated in Absolut!.
Conclusion: Generating high-affinity antibodies appears feasible with only a few hundred affinity measurements.
See full paper published in Cell Reports Methods:
Toward real-world automated antibody design with combinatorial Bayesian optimization
3) Designing antibodies robust to escape mutations via opponent shaping
Goal: Engineer antibodies that steer viral evolution toward controlled variants rather than escape.
Approach: Simulate evolutionary dynamics with Absolut! and compare design strategies; identify regimes that keep the virus in check.
See full paper published ICML 2025: ADIOS: Antibody Development via Opponent Shaping

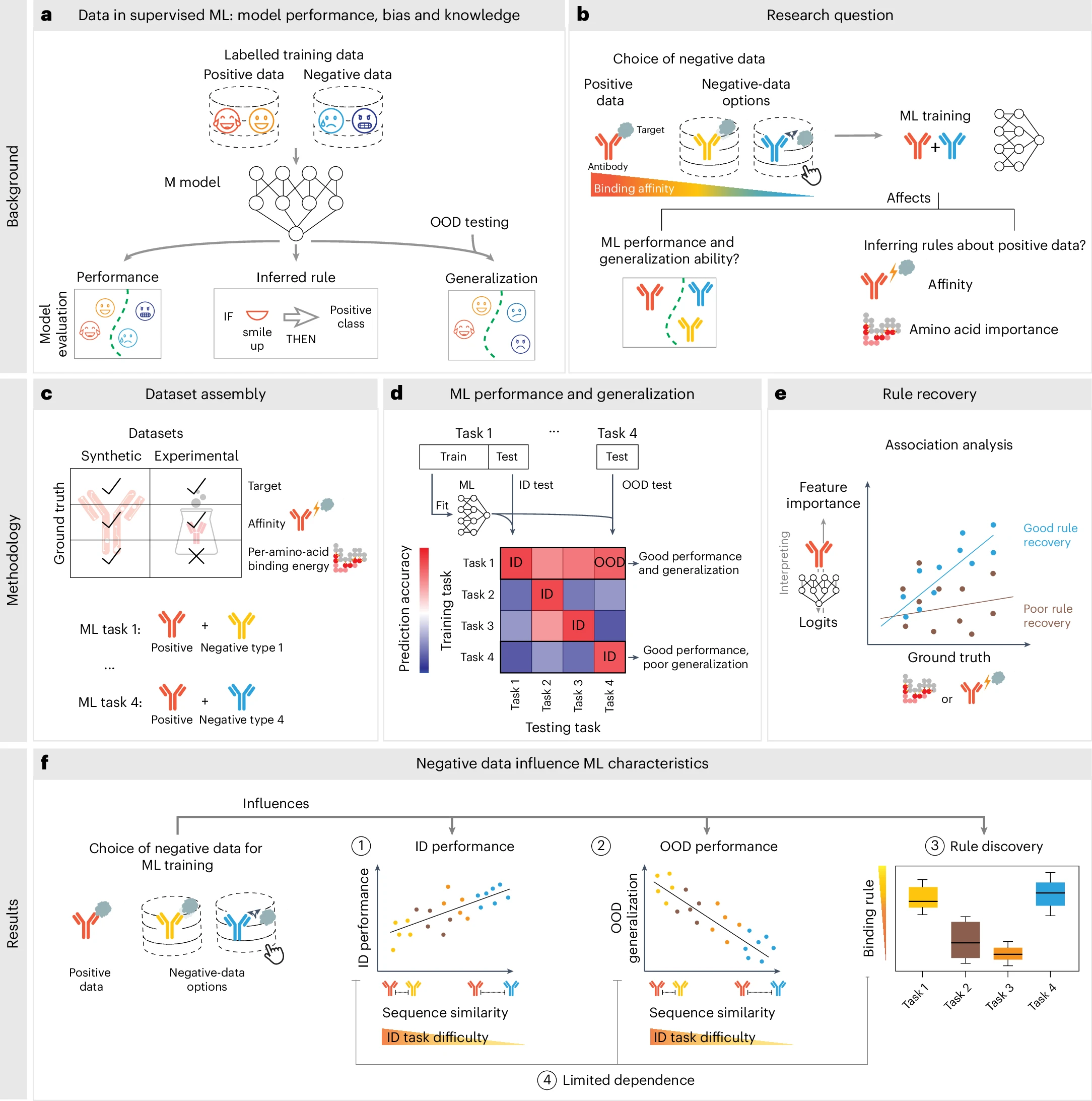
4) Define positive and negative datasets to improve ML generalizability
Finding: Pose-ranking models improve when negatives are more stringently defined. This was reproduced on Absolut! and translates to small-molecule binding.
Notes: Data enhancement and diverse non-binding pairs improved performance tested on experimental datasets.
See full paper published Nature Machine Intelligence:
Training data composition determines machine learning generalization and biological rule discovery